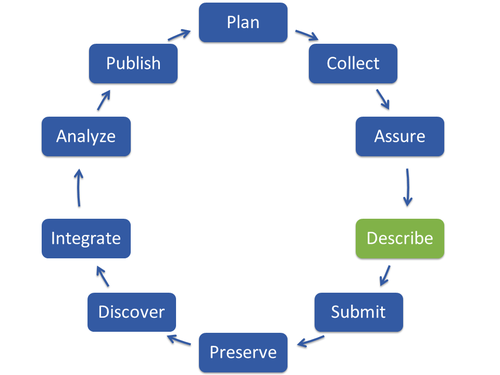
Describe
What is it?
After assuring the quality of your data, a detailed description of the data set should be provided, so other users can easily find your data, understand the context and content, reuse and cite your data. Your published article may not be sufficient to gain this information. The description of data generates structured information, so-called metadata, which should answer the following questions:
WHY were the data generated? WHO created the data? WHERE and WHEN were the data collected? WHAT is the content of the data? HOW were the data assessed?
How to do it?
- To save time, start early with the description of your data – as long as the information is present.
- Define how you want your data to be cited.
- Use appropriate metadata standards where possible to avoid errors and to make your metadata compatible.
- The description should include the technical context (names of datasets and data files in the datasets, versioning, file format, (processing of data), hardware/software, methods, tools, instruments used for data collection.
- State who was and is involved in the study (collectors, stakeholders, funders, contact person for questions).
- Describe the scientific context: Why were the data collected (hypothesis)? What kind of data were collected? Where were the data collected and when? Which standards or calibrations were used?
- State the precision with which each parameter was generated/measured. What are the units and formats? Which codes and abbreviations were used?
- Use the GFBio-Terminology-Service to avoid the usage of different names for e.g. the same species. By accounting for synonyms and acronyms, terms are kept consistent and reliable and your data can be annotated.
Who does it?
Ideally everybody who is producing data.
Key Elements
- Write metadata in a way that your data can be understood by other persons without additional knowledge - be precise, don’t use abbreviations or jargon, ideally use a metadata standard.
- Alternatively, convert your metadata afterwards to a compatible standard (e.g. EML, ABCD).
- Make sure the 6 questions can be answered (Who? How? Where? When? What? Why?)
GFBio Services
Data Curation
- Support for mapping your metadata to a compatible standard
Terminology Service (TS)
- GFBio Terminology Service: create, publish and curate your own terminologies or use existing terminologies
Useful Links
https://www.dataone.org/best-practices (Data One Best practice)
http://www.dcc.ac.uk/sites/default/files/documents/metadata%5B1%5D.pdf (DCC)
http://www.youtube.com/watch?v=7IN_SD5B43U (MANTRA Video with Lynn Jamieson)
http://rd-alliance.github.io/metadata-directory/ (overview of DCC metadata standards)
https://www.youtube.com/watch?v=-MIH8PkuUo4&feature=relmfu (a data file called SAM)
Recommended citation:
German Federation for Biological Data (2021). GFBio Training Materials: Data Life Cycle Fact-Sheet: Data Life Cycle: Describe. Retrieved 16 Dec 2021 from https://www.gfbio.org/training/materials/data-lifecycle/describe.